 Department of Computer Science
Department of Computer Science
Computational selection of transcriptomics experiments improves Guilt-by-Association analyses
Prajwal Bhat, Haixuan Yang, László Bögre, Alessandra Devoto and Alberto Paccanaro
PLoS ONE, vol. 7, iss. 8, p. 39681, 2012. 
Microarray is perhaps the most widely used gene expression measurement technology today. Gene expression profiles obtained from microarray experiments provide an insight into gene function and it is considered a powerful exploratory technique. However, understanding gene function from the expression profiles is a challenge. One of the widely used principles for elucidating the function of genes is the Guilt by Association (GBA) principle where, genes are considered to be involved in similar functions if the gene expression profiles are correlated. GBA principle is used in a variety of functional analyses techniques ranging from simple co-expression analysis, clustering techniques to network based approaches.
Generally in gene functional analyses, the GBA principle is applied to large collections of microarrays from various biological backgrounds such as different treatments and tissues. However, it is generally accepted that using large heterogeneous collections of microarrays could sometimes be unreasonable due to the condition specific nature of gene expression. This is evident when we observe the distribution of correlation coefficients among genes which are involved in the same function (See figure 1A below). If genes in the same functional category show poor correlation, then based on correlation, identifying new genes which might be involved in that function would be not possible.
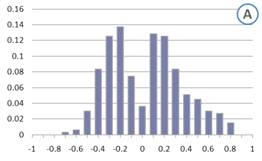
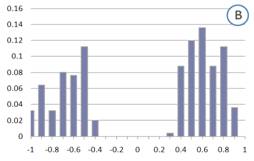
Figure 1: Distribution of correlation coefficients for genes in the GO category GO:009753 “Response to Jasmonic Acid stimulus” calculated using (a) a large heterogeneous collection of experiments and (b) a manually selected set of experiments, which were deemed to be functionally relevant to Jasmonic Acid response based on literature knowledge. To account for the different vector length in the two cases, correlation coefficients were counted only when significant (p-value<0.05).
In our work, we have shown that using only those experiments which are relevant to the functional category of interest could greatly improve the correlation between the genes in the same functional category (figure 1B). An obvious way to identify experiments that may be relevant to the analysis is by simply reading the literature. However, with ever increasing size of microarray compendia, identifying relevant experiments by literature review alone is an impractical task. In our paper, we present an algorithm that identifies functionally relevant experiments from a large collection of microarrays.
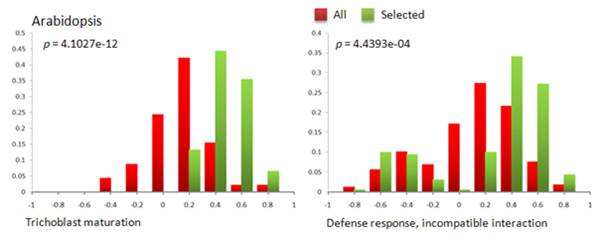
Figure 3: Distribution of correlation coefficient in GO BP terms show that the selected set of experiments enrich higher correlations when compared to using all experiments. The p-values from the t-test indicate that the distribution of correlations for the selected set is significantly greater than for all experiments.
We find that the experiments selected by our algorithm lead to a significantly higher distribution of correlation between the genes which belong to the same functional category when compared to using all experiments in the compendia (See Figure 2). We also find that a GBA based classifier will consistently outperform when experiments selected by our algorithm are used for calculating the correlation compared to using all experiments. Interestingly, the performance of the GBA based classifier when selected experiments are used varies with specificity of the functional annotation i.e. the performance is better with the more specific functional categories (lower nodes of the GO Biological process tree) compared to the broader functional categories (higher nodes in the GO tree).
The MATLAB code for the algorithm and the data used in the paper can be downloaded from here

